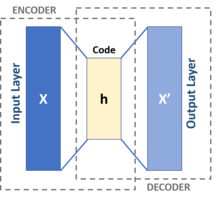
Comparative Study of Deep Generative Models on Chemical Space Coverage | Journal of Chemical Information and Modeling
Partitioning variability in animal behavioral videos using semi-supervised variational autoencoders | PLOS Computational Biology

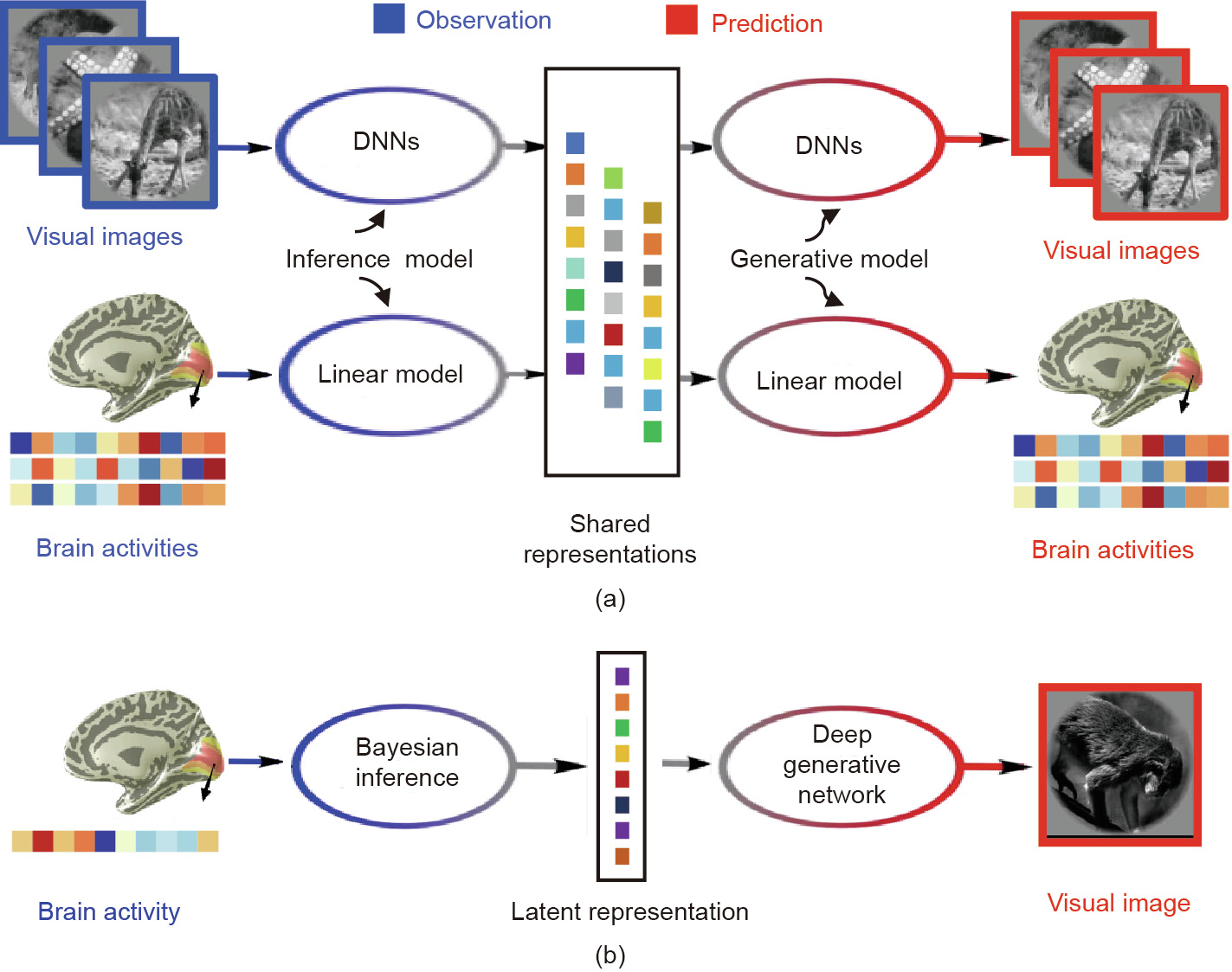
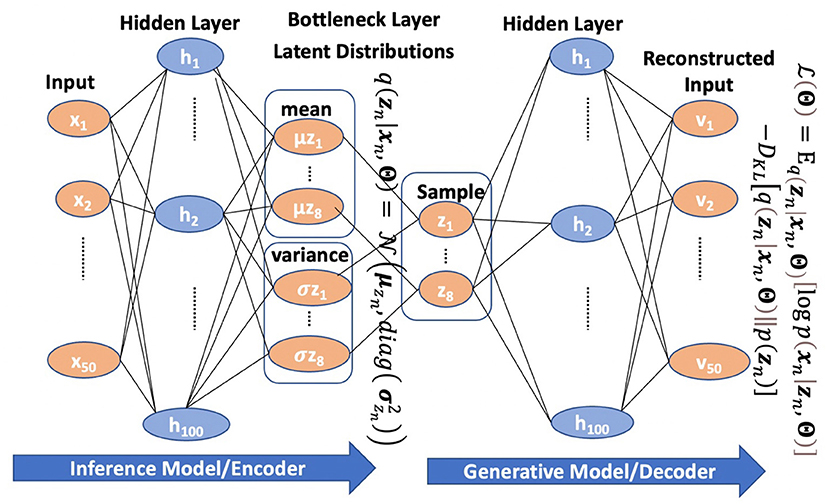
Variational Autoencoder: An Unsupervised Model for Modeling and Decoding fMRI Activity in Visual Cortex | bioRxiv

Variational autoencoder: An unsupervised model for encoding and decoding fMRI activity in visual cortex. - Abstract - Europe PMC
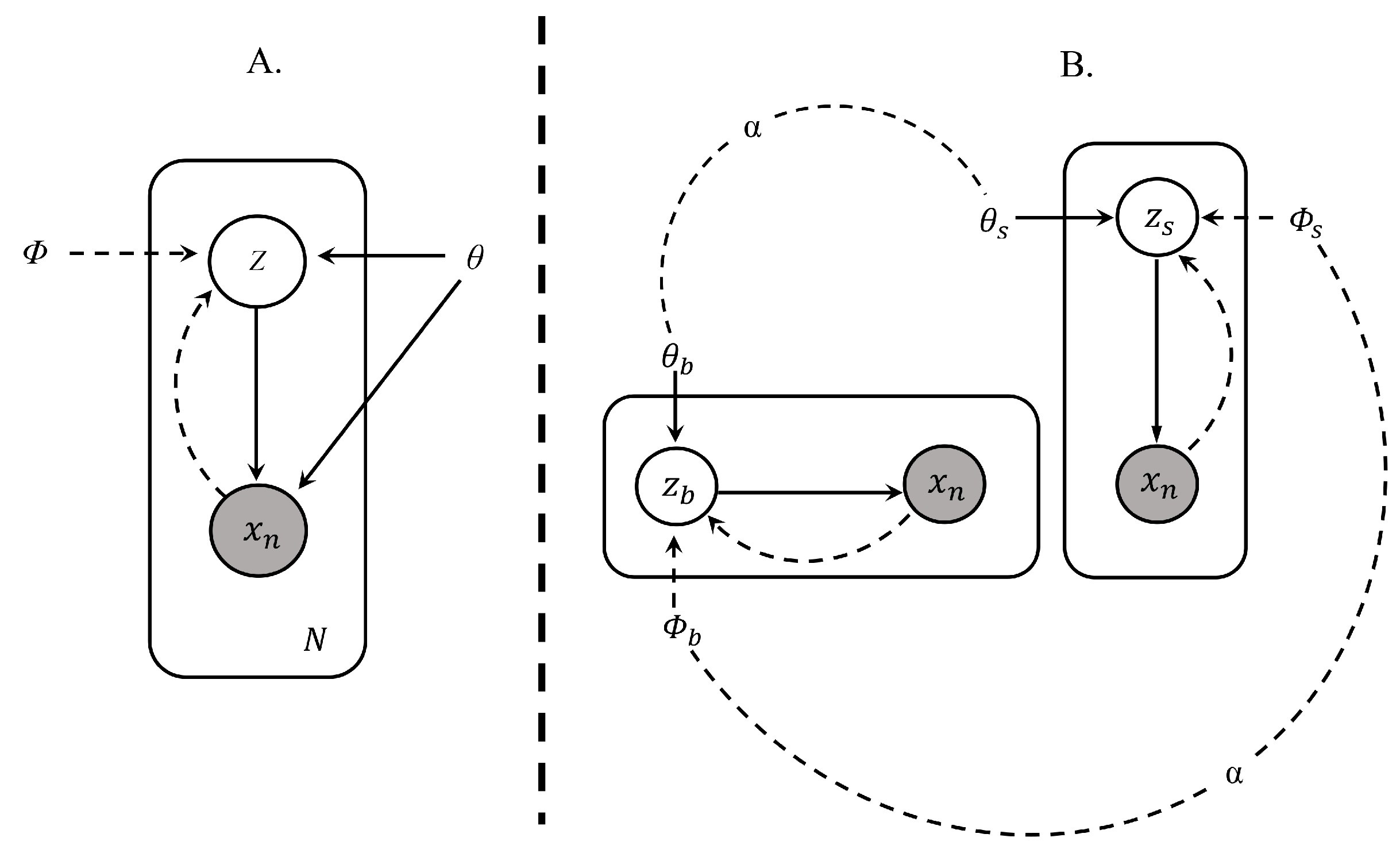
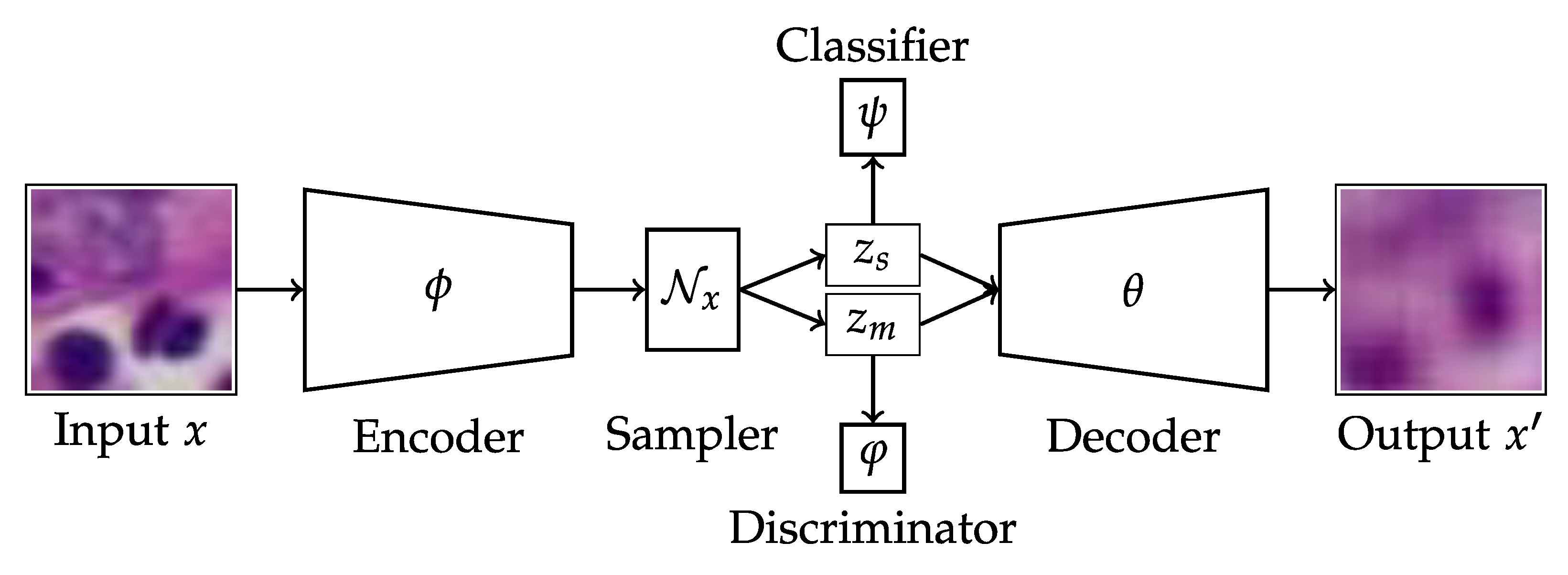
Applied Sciences | Free Full-Text | Improving Generative and Discriminative Modelling Performance by Implementing Learning Constraints in Encapsulated Variational Autoencoders | HTML
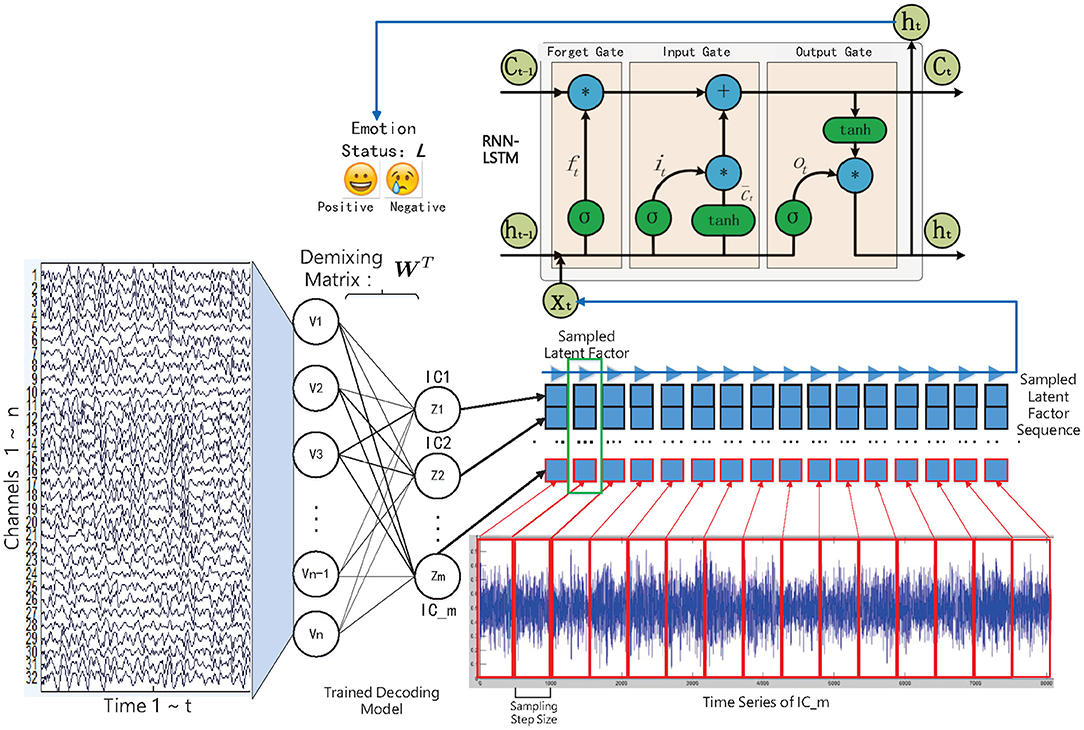
Frontiers | Latent Factor Decoding of Multi-Channel EEG for Emotion Recognition Through Autoencoder-Like Neural Networks

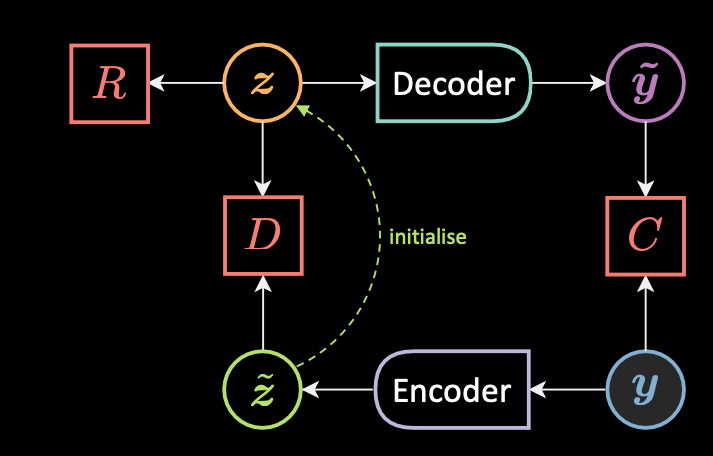
Molecular Generative Model Based on an Adversarially Regularized Autoencoder | Journal of Chemical Information and Modeling

De Novo Molecule Design Using Molecular Generative Models Constrained by Ligand–Protein Interactions | Journal of Chemical Information and Modeling

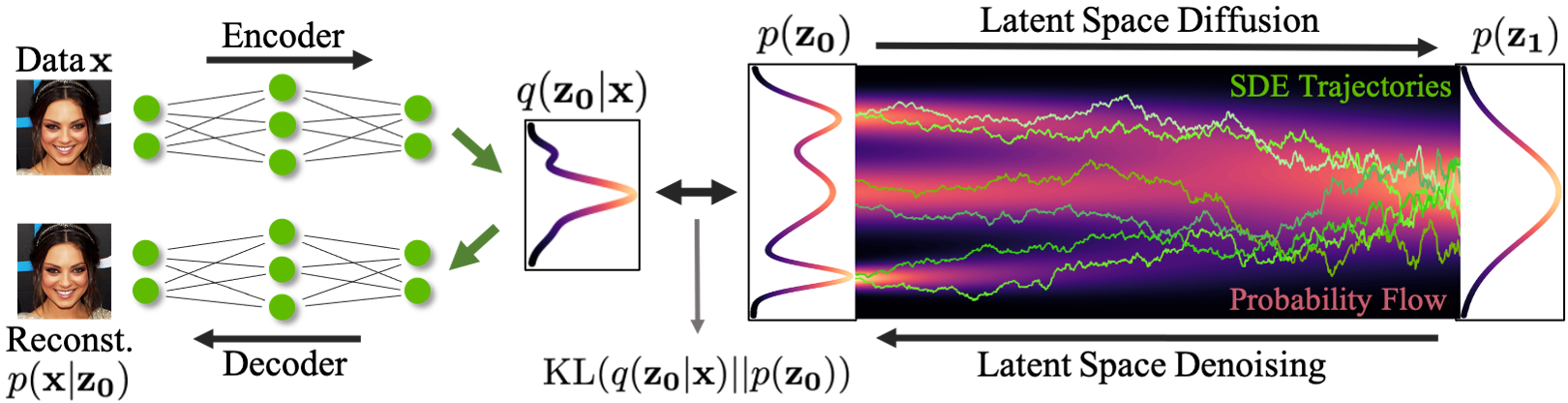
Enhancing scientific discoveries in molecular biology with deep generative models | Molecular Systems Biology

Reconstructing feedback representations in ventral visual pathway with a generative adversarial autoencoder | bioRxiv

Molecular Generative Model Based on an Adversarially Regularized Autoencoder | Journal of Chemical Information and Modeling
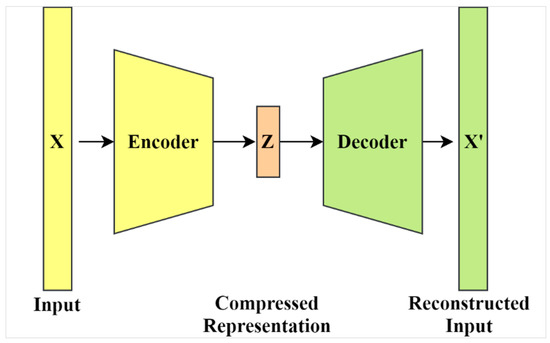
J. Imaging | Free Full-Text | Variational Autoencoder for Image-Based Augmentation of Eye-Tracking Data | HTML
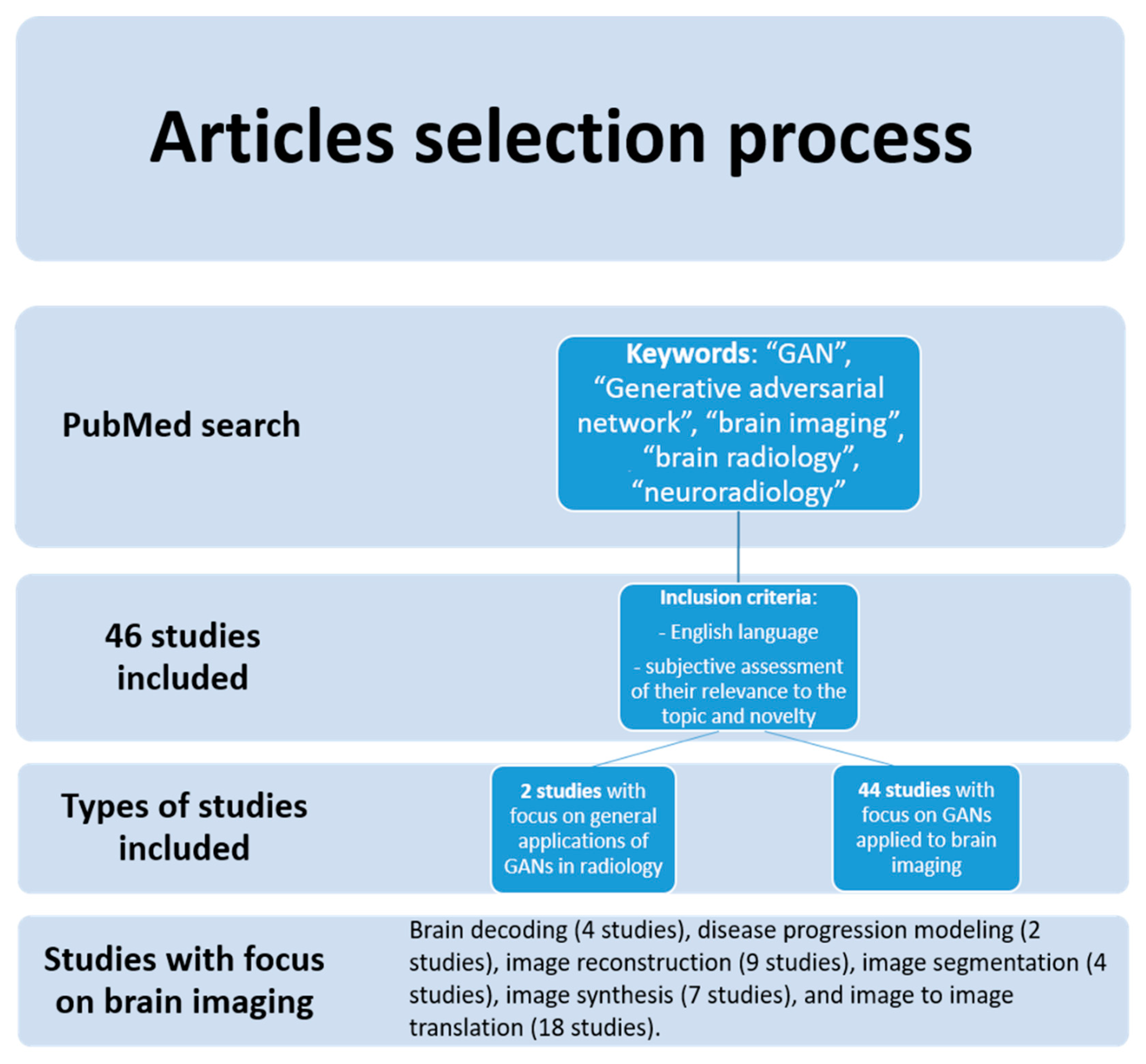
J. Imaging | Free Full-Text | Generative Adversarial Networks in Brain Imaging: A Narrative Review | HTML