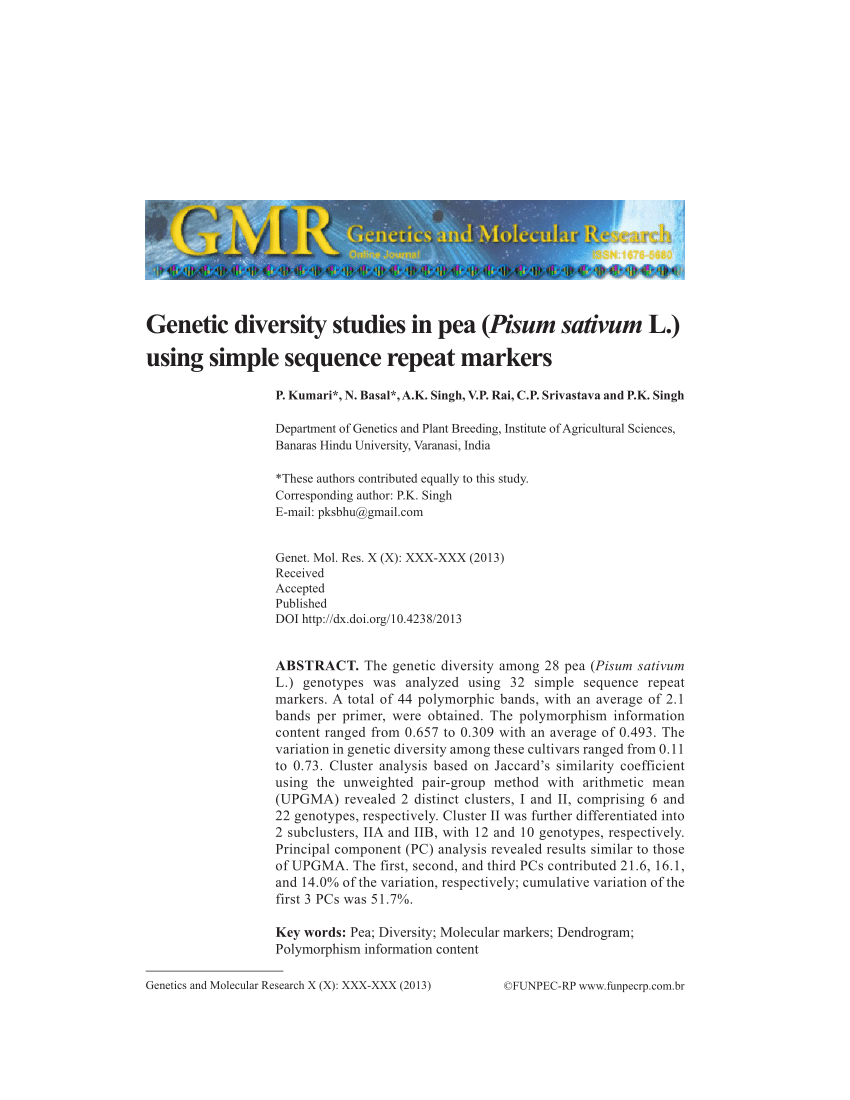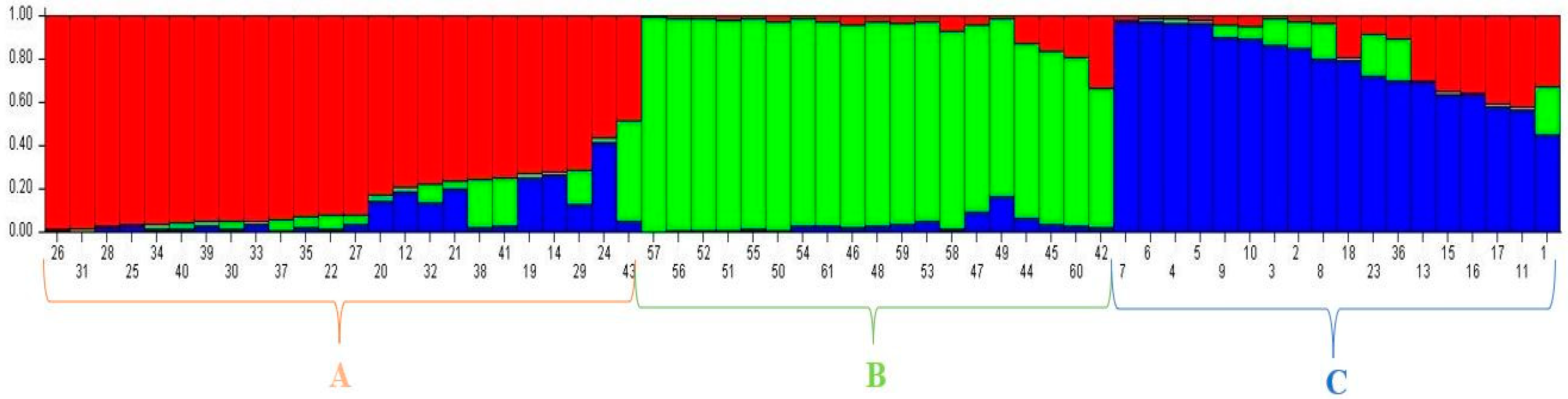
Genes | Free Full-Text | SSR-Based Molecular Identification and Population Structure Analysis for Forage Pea (Pisum sativum var. arvense L.) Landraces | HTML

A community resource for exploring and utilizing genetic diversity in the USDA pea single plant plus collection | Horticulture Research

PDF) Genetic Diversity and Population Structure Among Pea (Pisum sativum L.) Cultivars as Revealed by Simple Sequence Repeat and Novel Genic Markers
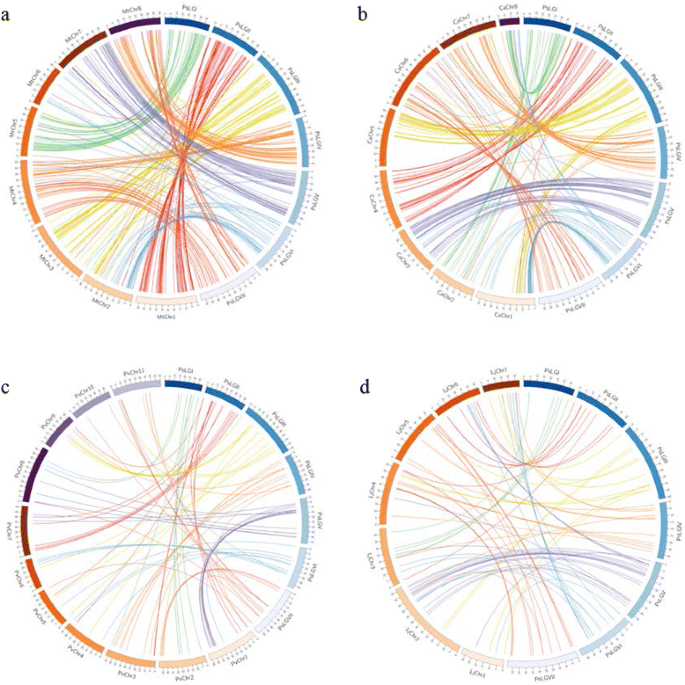
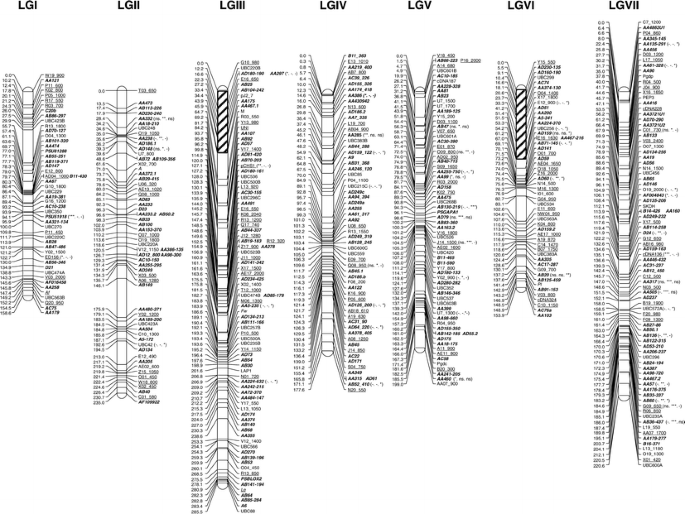
Genome-wide SNP identification, linkage map construction and QTL mapping for seed mineral concentrations and contents in pea (Pisum sativum L.) | BMC Plant Biology | Full Text

Comprehensive functional analysis and mapping of SSR markers in the chickpea genome (Cicer arietinum L.) - ScienceDirect

PDF) Estimation of pea ( Pisum sativum L.) microsatellite mutation rate based on pedigree and single-seed descent analyses | Petr Smykal - Academia.edu
Pea Marker Database (PMD) – A new online database combining known pea (Pisum sativum L.) gene-based markers | PLOS ONE

Genome-wide SNP identification, linkage map construction and QTL mapping for seed mineral concentrations and contents in pea (Pisum sativum L.) – topic of research paper in Biological sciences. Download scholarly article PDF
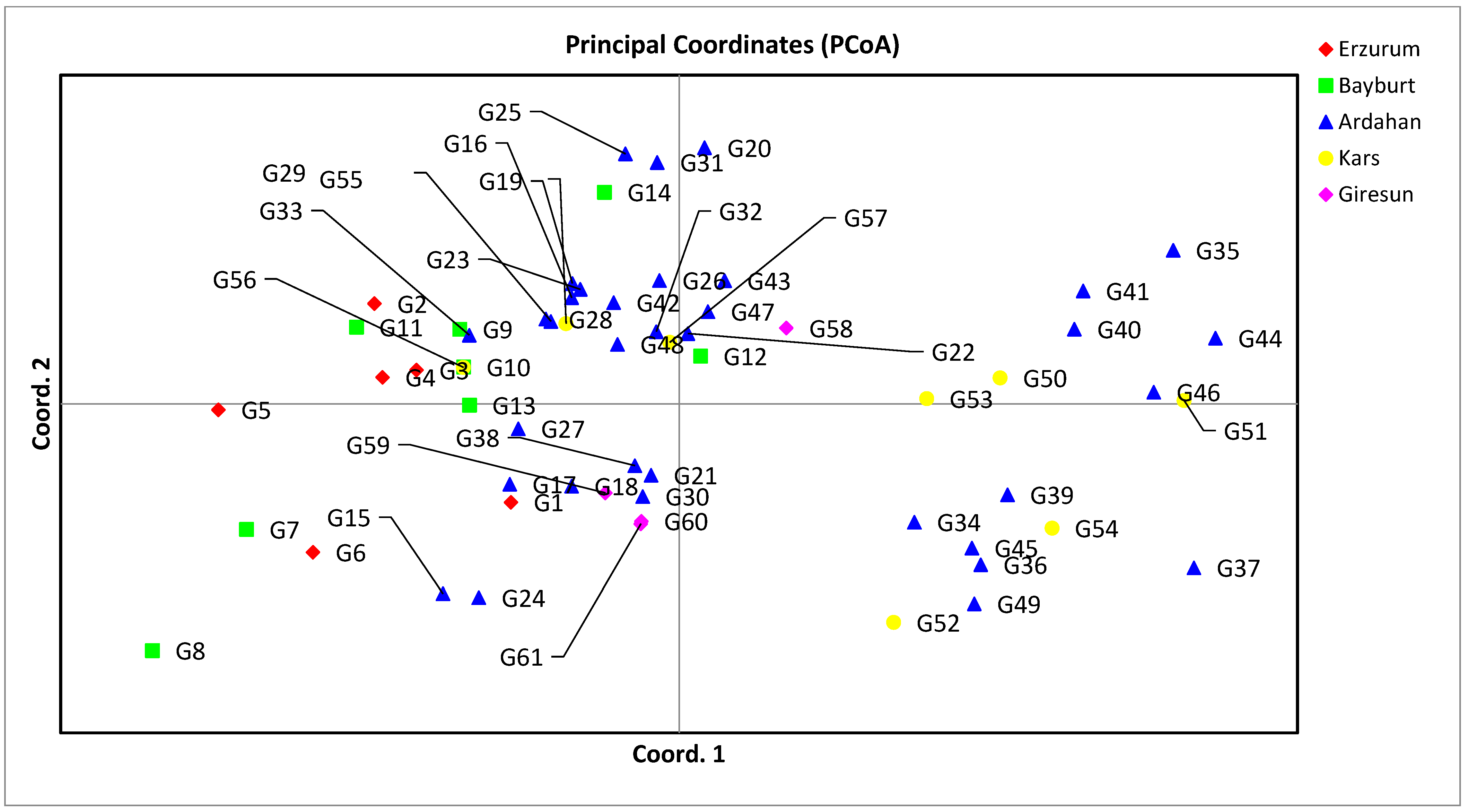
Genes | Free Full-Text | SSR-Based Molecular Identification and Population Structure Analysis for Forage Pea (Pisum sativum var. arvense L.) Landraces | HTML
High-Throughput Development of SSR Markers from Pea (Pisum sativum L.) Based on Next Generation Sequencing of a Purified Chinese Commercial Variety | PLOS ONE
![PDF] Molecular characterization of pea (Pisum sativum L.) using microsatellite markers | Semantic Scholar PDF] Molecular characterization of pea (Pisum sativum L.) using microsatellite markers | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6b50c024ad7293ae031ff7e23da4eb0630955b03/3-Table2-1.png)
PDF] Molecular characterization of pea (Pisum sativum L.) using microsatellite markers | Semantic Scholar

Genome-wide SNP identification, linkage map construction and QTL mapping for seed mineral concentrations and contents in pea (Pisum sativum L.) | BMC Plant Biology | Full Text
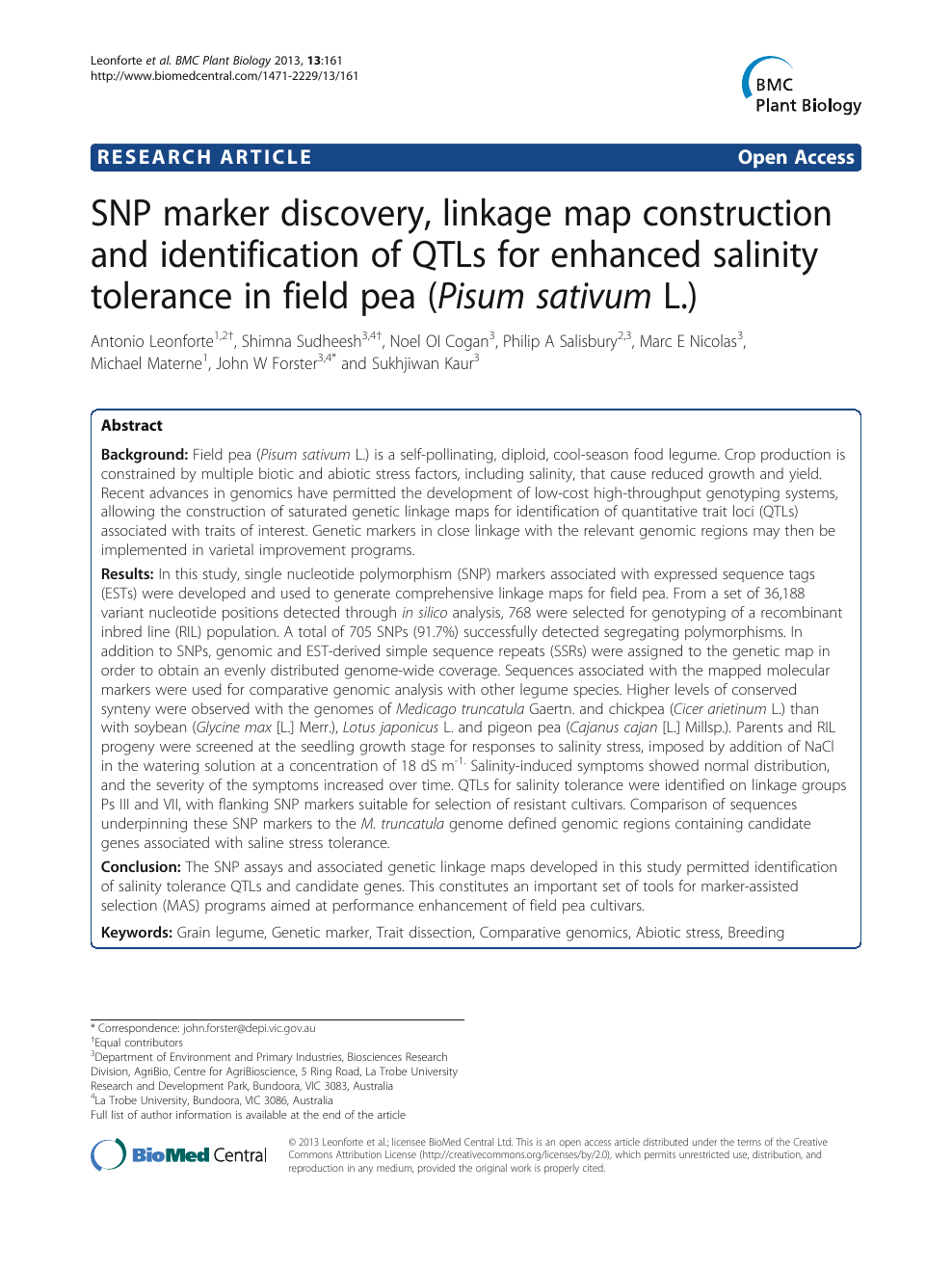

SNP marker discovery, linkage map construction and identification of QTLs for enhanced salinity tolerance in field pea (Pisum sativum L.) – topic of research paper in Biological sciences. Download scholarly article PDF
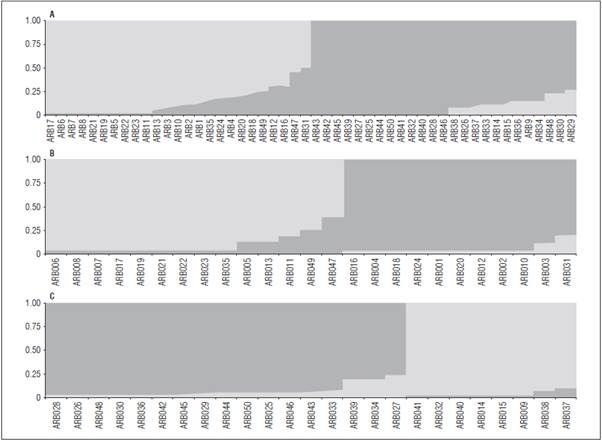
Molecular characterization using SSR markers in 50 shrub pea genotypes (Pisum sativum L.) from the GRICAND Collection, Colombia
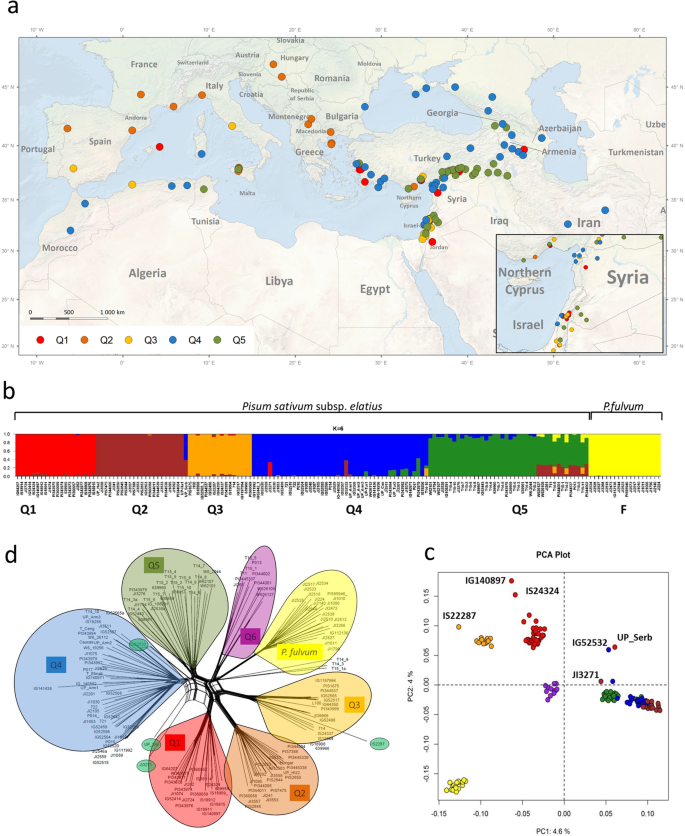
Genomic diversity and macroecology of the crop wild relatives of domesticated pea | Scientific Reports

PDF) Genetic diversity in pea (Pisum sativum L.) accessions detected by sequence tagged microsatellite markers

Development of EST‐derived SSR markers in pea (Pisum sativum) and their potential utility for genetic mapping and transferability - Mishra - 2012 - Plant Breeding - Wiley Online Library